NGDC Dec 17, 2020
Genome sequence variation is a kind of heritable variation occurring on DNA level. It is one of the most valuable genetic data resources for studies such as species population genetic evolution, phenotypic difference, human disease research, animal and plant molecular breeding. In recent years, with the development of sequencing technology, more and more genomes have been analyzed precisely, therefore the variation data of whole genome sequences from different species and different populations are growing rapidly.
In order to facilitate the open sharing and security management of the scientific data of variation genomics from different biological genetic resources, the Genome Variation Map (GVM), a public data repository of genome variations, was launched in 2017 by the researchers of the National Genomics Data Center, Beijing Institute of Genomics of Chinese Academy of Sciences/ China National Center for Bioinformation (CNCB), and has been growing as the largest sequence variation database of multi species genomes in China. Recently, the GVM version 2.0 was successfully updated. This work was published online on November 10 in Nuclear Acids Research with the title "Genome Variation Map: a worldwide collection of genome variations across multiple species".
Compared with the previous version, particularly, a total of 22 species, 115 projects, 55,935 samples, 463,429,609 variants, 66,220 associations and 56 submissions were newly added in the current version of GVM. In the current release, GVM houses a total of ~960 million variants from 41 species, including 13 animals, 25 plants and 3 viruses. Moreover, it incorporates 64,819 individual genotypes and 260,393 manually curated high quality genotype-to-phenotype associations. Since its inception, GVM has archived genomic variation data of 43,754 samples submitted by worldwide users and served >1 million data download requests. Collectively, as a core resource in the National Genomics Data Center, GVM provides valuable genome variations for a diversity of species and thus plays an important role in both functional genomics studies and molecular breeding.
Through structural data organization, the GVM database has developed modules of user-friendly data searching, browsing, submission, download, and statistics. Users can easily and quickly browse the detailed information of stored species, projects, samples, variations, association knowledge and submitted data. Under the searching module, users can easily obtain all variation data of a species, with the functional knowledge information, annotation gene information, population frequency and so on. Users can also download the whole genome sequence variation data in VCF or FASTA format through FTP service.
With the active response to the requirements of scientific data management and sharing, the GVM database has established an online submission module of genome sequence variation data. It can provide batch data online delivery service, and assign a unique identifier for each submitted data. Data submitter is required to send application of data submission through personal account. The submitted data would be released according to submitter's requirement with controllable management in GVM. Relying on the high-performance storage and remote disaster recovery backup mechanism of CNCB, the GVM database system will keep data updated and backup regularly to ensure data integrity and security.
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences, National Key R&D Program of China, etc.
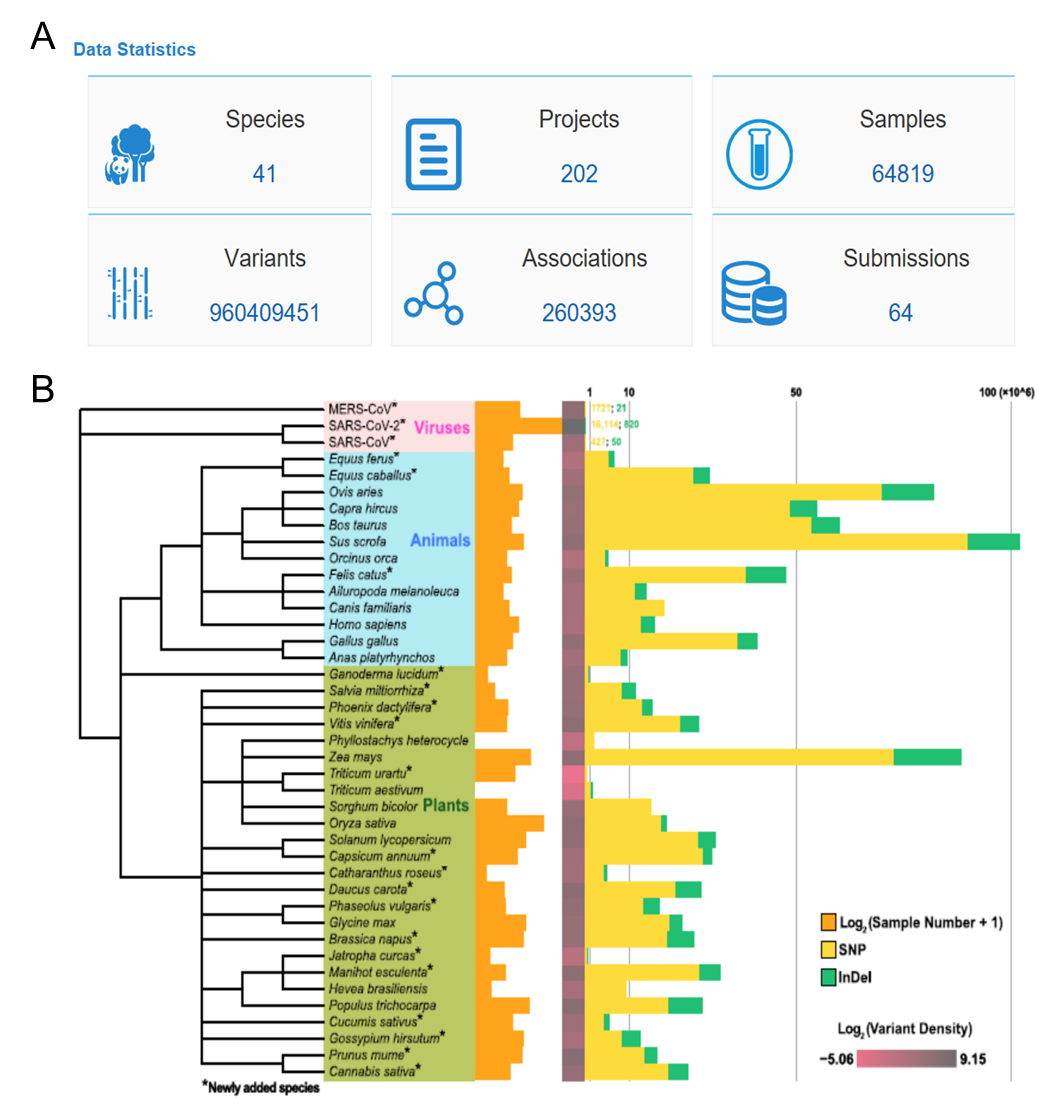
GVM database: (a) six functional modules and data volume; (b) statistical chart of variation data volume and density of each species (Image by NGDC)
Contact:
Dr. SONG Shuhui
E-mail: songshh@big.ac.cn
Dr. ZHANG Zhang
E-mail: zhangzhang@big.ac.cn