CAT (Composition Analysis Toolkit) is a software package that includes a novel measure of codon usage bias, Codon Deviation Coefficient (CDC). Unlike previous measures, CDC effectively accounts for background nucleotide composition in estimating codon usage bias and utilizes a bootstrap assessment of the statistical significance of codon usage bias.
Please cite: Zhang, Z., Li, J., Cui, P., Ding, F., Li, A., Townsend, J.P., and Yu, J. (2011) Codon Deviation Coefficient: a novel measure for estimating codon usage bias and its statistical significance, under review.
The CAT package, including source codes, compiled executables, and documentation, is freely available for academic use only.
For windows, linux and unix, the CAT package (version 1.3) can be downloaded here
CAT version 1.0 for MAC OS with Graphical User Interface (GUI) can be downloaded here. Below is a screenshot:
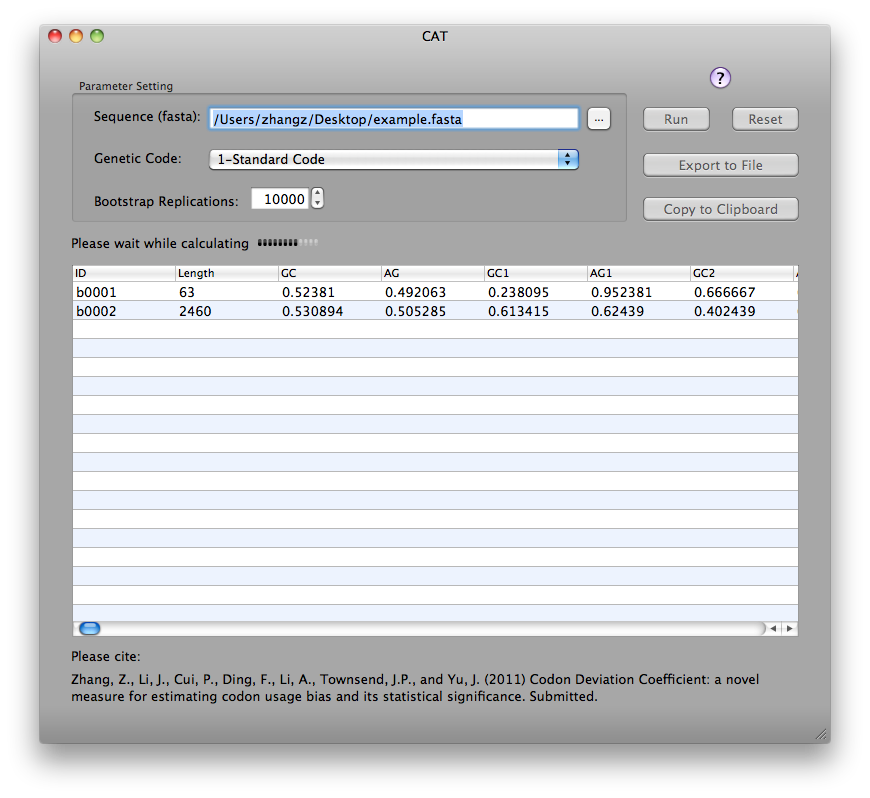
CAT is distributed as open-source software and licensed under the GNU General Public License (Version 3; http://www.gnu.org/licenses/gpl.txt), in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
Commercial use of CAT requires a special contract.
For high efficiency and compatibility with more platforms, CAT is written in standard C++. The package is normally named CATXXX.tar.gz (XXX stands for the version).
Executables have been precompiled for Linux/Unix/Mac/Windows. Please unpack the package of CATXXX.tar.gz (see below) and then you will find compiled executables in the folder of "CATXXX/bin/".
For compilation on your specific platform, please follow the steps below.
Unpack the package of CATXXX.tar.gz by the following commands.
tar -zxf CATXXX.tar.gz
If you use other Linux/Unix/Mac OS, you have to compile the program in the source codes folder with the help of g++/gcc compiler.
cd CATXXX/src
make
That's it. Then you can find an executable named "CAT" in this folder.
Note for Mac users: Mac on your computer might use the case insensitive file system, so that "CAT" would have the completely "same" name with a system command "cat". When running the "CAT" program, please specify the working directory of "CAT" for access.
CAT allows the user to customize parameters. The following are the parameters' settings, which can also be found by typing "CAT -h".
-i input fasta file name [string, required]-o output file name [string, optional], default = input file name with the characters ".cat"
appended-b bootstrap replications [integer, optional], default = 10000-c genetic code to be used [integer, optional], default = 1. All genetic codes are available
at NCBI.CAT accepts FASTA file (http://www.ncbi.nlm.nih.gov/BLAST/fasta.shtml) which contains multiple nucleotide coding sequences. Stop codons are eliminated from the analysis.
Example:
>b0001
ATGAAACGCATTAGCACCACCATTACCACCACCATCACCATTACCACAGGTAACGGTGCGGGCTGA
>b0075
ATGACTCACATCGTTCGCTTTATCGGTCTACTACTACTAAACGCATCTTCTTTGCGCGGTAGACGAGTGAGCGGCATCCAGCATTAA
>b1265
ATGAAAGCAATTTTCGTACTGAAAGGTTGGTGGCGCACTTCCTGA
An example data file as well as its results file accompanies the CAT package in the folder "CATXXX/example/".
CAT output is in the form of a tab-delimited text file with one header row. Each row thereafter displays the results for each single gene, including columns with gene ID and gene length (bp), GC and purine contents, the estimates of CDC and its significance level P-value. In addition, the observed and expected compositions of nucleotides, codons and amino acids are also provided.
The description for each column is listed as follows.
In addition, observed and expected compositions for nucleotide (3*4), codon (64) and amino acid (20) are also outputted.
The output file name, by default, will be same as the original input file name with the characters ".cat" appended. In addition, the output file name can also be customized by setting the parameter "-o output_filename". Please see details in the section of "Setting Parameters".
We thank Joe Yu for constructive comments on this work and George Marselis for providing assistance on web page hosting. We also thank many users for reporting bugs and sending suggestions.
Please send bugs or advice to Dr. Zhang Zhang (zhangzhang@big.ac.cn, zhangzhang.cn@gmail.com).