The 3rd Big Data Forum for Life and Health Sciences
The 3rd Big Data Forum for Life and Health Sciences (October 11-14, 2018)
Biological research has entered the era of big data, including a wide variety of omics data and covering a broad range of health data. Such big data is generated at ever-growing rates and distributed throughout the world with heterogeneous standards and diverse limited access capabilities. However, the promise to translate these big data into big knowledge can be realized only if they are publicly shared. Thus, providing open access to omics & health big data is essential for expedited translation of big data into big knowledge and is becoming increasingly vital in advancing scientific research and promoting human healthcare and precise medical treatment.
Open Biodiversity & Health Big Data
It is our great pleasure to announce that the 2018 Big Data Forum for Life and Health Sciences will be held in October 11-14, 2018. A few renowned biomedical data scientists have agreed to give speeches. Likely, you are also cordially invited to share your work and participate in this excited event.
Looking forward to seeing you in Beijing, China! We will be working hard to ensure your stay not only a fruitful one, but also an enjoyable one!
会议通知下载
Organizing Committee
- Yiming Bao (BIG, CAS)
- Zhang Zhang (BIG, CAS)
- Wenming Zhao (BIG, CAS)
- Jingfa Xiao (BIG, CAS)
- Songnian Hu (BIG, CAS)
- Jun Yu (BIG, CAS)
- Jingchu Luo (Peking University)
Registration

Previous Conferences

Invited Speakers

Amir Abbasi
Professor
National Centre for Bio Informatics
Quaid-i-Azam University
Pakistan

Yiming Bao
Professor
BIG Data Center, Beijing Institute of Genomics
Chinese Academy of Sciences
China

Suhua Chang
Associate Professor
National Clinical Research Center for Mental Disorders
Institute of Mental Health, Peking University
China

Kaifu Chen
Associate Professor
Center Director for Bioinformatics and Computational Biology
Houston Methodist, Weill Cornell Medical College
USA

Luonan Chen
Professor
Shanghai Institute of Biochemistry and Cell Biology
Chinese Academy of Sciences
China

Frank Eisenhaber
Executive Director
Head of Division, Biomolecular Function Discovery
Bioinformatics Institute
Singapore

Michael Y. Galperin
Lead Scientist
Computational Biology Branch
National Center for Biotechnology Information
USA

Zhiyuan Hu
Professor
National Center for Nanoscience and Technology
Chinese Academy of Sciences
China

Yuxia Jiao
Editor of Genomics, Proteomics & Bioinformatics
Beijing Institute of Genomics, CAS
China

Cheng Li
Professor
School of Life Sciences
Peking University
China

Mengwei Li
PhD Candidate
BIG Data Center, Beijing Institute of Genomics
Chinese Academy of Sciences
China

Xia Li
Professor
College of Bioinformatics Science and Technology
Harbin Medical University
China

Guoqing Lu
Isaacson Professor, Genomics and Bioinformatics
Department of Biology & School of Interdisciplinary Informatics
University of Nebraska at Omaha
USA

Hui Lu
Professor
School of Life Sciences and Biotechnology
Shanghai Jiao Tong University
China

Vsevolod J. Makeev
Professor
Dept. Computational Systems Biology
Vavilov Institute of General Genetics, RAS
Russia

Suchinda Malaivijitnond
Professor
Director, National Primate Research Center of Thailand
Chulalongkorn University
Thailand

Daniel Stekhoven
Head
Clinical Bioinformatics Unit
ETH Zurich
Switzerland

Zhixi Su
Professor
School of Life Sciences
Fudan University
China

Qianfei Wang
Professor
Associate Director of Key Laboratory of Genomic and Precision Medicine
Beijing Institute of Genomics, CAS
China

Xiangfeng Wang
Professor
Director of Department of Crop Genomics and Bioinformatics
College of Agronomy and Biotechnology, China Agricultural University
China

Xiyin Wang
Professor
College of Life Sciences
North China University of Science and Technology
China

Changqing Zeng
Professor
CAS Key Laboratory of Genomic and Precision Medicine
Beijing Institute of Genomics, CAS
China

Xiaojun Zhang
Professor
Experimental Marine Biological Laboratory
Institute of Oceanology, CAS
China

Hongkun Zheng
CEO
Biomarker Technologies
China

Qing Zhou
Professor
Life Sciences Institute
Zhejiang University
China
Agenda
| October 11: Pick-up & Registration | |
|---|---|
| October 12: Talks | |
| 09:00 - 10:10 | Session 1, chaired by Yiming Bao, BIG, CAS |
| 09:00 - 09:10 | Welcome and Opening Remarks
Zhang Zhang, On Behalf of the Organizing Committee, BIG, CAS |
| 09:10 - 09:40 | Phase 1 of CASPMI Project and Data Analysis [Abstract]
Changqing Zeng, BIG, CAS
Launched by the Chinese Academy of Sciences (CAS) in 2016, the Phase 1 of the CAS Precision Medicine Initiative Project (CASPMI) aims at (i) next generation sequencing of the whole genome (25-30X) for 1000 samples collected in CAS cohort; (ii) construction of a reference genome from a northern Han individual (NH1.0) using a hybrid approach including PacBio sequencing,
10X Genomics library preparation, and Bionano optical mapping; (iii) construction of electronic health records and genetic reports for CASPMI participants; (iv) association analyses based on sequencing data and phenotypes obtained from base line collection of the project. I will introduce and summarize the current results of CASPMI project. In brief, near the completion of the phase 1, we are able to provide a comprehensive genetic variation map including 24.85M SNPs, 3.85M small indels and 106,382 structural variations.
In total, we identified population-specific variations of 55,271 SNPs and 6,774 indels in this cohort study, among which 42 significant SNPs in 39 genes are detected to present a significant correlation with various metabolic related traits and diseases based on GWAS-Catalog annotation.
Geographic differentiation of northern and southern populations was observed, as well as the mutational signatures of novel variants showed difference in these two groups. Variations in MTHFR, TCN2, FADS1, and FADS2, which are associated with circulating folate and vitamin B12 or lipid metabolism, suggest the selection from various environmental exposures and life styles especially dieting between northerners and southerners. The high-quality human genome assembly and a comprehensive genetic map will provide population-specific genetic variations for later studies of precision medicine and individualized healthcare.
|
| 09:40 - 10:10 | Human Brain Evolution In Context of Enhancer Divergence [Abstract]
Amir Abbasi, Quaid-i-Azam University, Pakistan
Humans are usually considered to be far the most intelligent than others animals. Factors that make up the basis of brain properties include size of the brain, cortex, prefrontal cortex and degree of encephalization. Now the question is how we can interpret the phenomenal complexity of Human Brain? The sophistication of vertebrate Brain is orchestrated through the signalling cascade of cis-regulatory modules, so to decipher this signal co-ordination is obligatory to comprehend prototyping of brain.
In this talk we will be focusing on the acceleration in non-coding regulatory landscape of the genome and we will highlight the functional parts within it to have undergone accelerated divergence in present-day Human population. Moving ahead, we will focus on the transcription factors which are occupying the H. sapiens-unique binding sites such as SOX2 and RUNX1/3 and also play their part in maintaining a vital role in gene expression especially in the context of neural development.
The second objective of this talk will be to define the forebrain specific transcriptional code through which we predicted the 25000 Human forebrain specific enhancers on the basis of heterotypic clustering of the core transcription factors shortlisted through the code. These enhancers are now being validated through different strategies followed by their functional testing in Zebrafish.
|
| 10:10 - 10:40 | Group Photo and Tea & Coffee Break |
| 10:40 - 12:10 | Session 2, chaired by Jingfa Xiao, BIG, CAS |
| 10:40 - 11:10 | Utilizing Big Epigenomic Data for Cancer Gene Discovery [Abstract]
Kaifu Chen, Weill Cornell Medical College, USA [Personal Profile]
Kaifu Chen, PhD, is an Association Professor and is the Director for the Center For Bioinformatics and Computational Biology in the Methodist Hospital Research Institute and Cornell University Weil Cornell Medical College. His major research interest is to understand the epigenetic regulation of cancer development through bioinformatics interpretation of epigenome, genome, and transcriptome data.
Genes suspected of increasing the selective growth advantage of tumor cells were categorized as either Mut-driver genes or Epi-driver genes.
Recent genome sequencing efforts successfully detected millions of cancer mutations. However, it remains a challenge to define the catalogue of cancer driver genes by mutation analysis alone.
Only a small fraction of mutations in cancer actually affects driver genes. Meanwhile, many genes that do not mutate are epigenetically altered to drive cancer development. Unlike genetic sequence, epigenetic modifications vary with normal cell type, developmental stage, and biological environment.
Criteria have not yet been formulated for distinguishing epigenetic changes that exert a selective growth advantage from those that do not. We approach this challenge by integrating over 10,000 genomes and epigenomes to investigate epigenetic mechanisms that regulate cancer driver genes, and through novel bioinformatics strategy developed for cancer gene discovery using epigenetic signatures associated with these mechanisms.
Our research addresses the fundamental problem of how to identify cancer driver genes that are not mutated, but epigenetically altered in cancers to increase the selective growth advantage of tumor cells.
|
| 11:10 - 11:40 | Big data medicine by network biomarkers and dynamic network biomarkers
Luonan Chen, Shanghai Institute of Biochemistry and Cell Biology, CAS |
| 11:40 - 12:10 | P4 Medicine Journey in China [Abstract]
Zhiyuan Hu, National Center for Nanoscience and Technology, CAS
Systems medicine has united genomics and genetics through family genomics to more readily identify disease genes.
It has made blood a window into health and disease.
It is leading to the stratification of diseases (division into discrete subtypes) for proper impedance match against drugs and the stratification of patients into subgroups that respond to environmental challenges in a similar manner (e.g. response to drugs, response to toxins, etc.).
The convergence of patient-activated social networks, big data and their analytics, and systems medicine has led to a P4 medicine that is predictive, preventive, personalized, and participatory. Medicine will focus on each individual. It will become proactive in nature.
It will increasingly focus on wellness rather than disease. A journey has started to use P4 medicine strategy to prevent chronic diseases in China.
|
| 12:10 - 13:30 | Lunch and BIG tour |
| 13:30 - 15:10 | Session 3, chaired by Cheng Li, Peking University |
| 13:30 - 14:10 | Variant interpretation - how to tackle the bottleneck of comprehensive cancer diagnostics Daniel Stekhoven, ETH Zurich, Switzerland |
| 14:10 - 14:40 | Aberrant tRNA processing causes an autoinflammatory syndrome responsive to TNF inhibitors [Abstract]
Qing Zhou, Zhejiang University
We identified eight mutations in these nine patients, three of which have not been previously associated with SIFD.
Three patients died in early childhood. Inflammatory cytokines, mainly interleukin (IL)-6, interferon gamma (IFN-γ) and IFN-induced cytokines were elevated in the serum, whereas tumour necrosis factor (TNF) and IL-1β were present in tissue biopsies of patients with active inflammatory disease.
Deep tRNA sequencing of patients' fibroblasts showed significant deficiency of mature cytosolic tRNAs. EM of bone marrow and skin biopsy samples revealed striking abnormalities across all cell types and a mix of necrotic and normal-appearing cells. By immunoprecipitation, we found evidence for dysregulation in protein clearance pathways.
In 4/4 patients, treatment with a TNF inhibitor suppressed inflammation, reduced the need for blood transfusions and improved growth. Mutations of TRNT1 lead to a severe and often fatal syndrome, linking protein homeostasis and autoinflammation. Molecular diagnosis in early life will be crucial for initiating anti-TNF therapy, which might prevent some of the severe disease consequences.
|
| 14:40 - 15:10 | Biomedical Big Data to Knowledge --- Computational Systems Biology for Diseases
Xia Li, Harbin Medical University |
| 15:10 - 15:30 | Tea & Coffee Break |
| 15:30 - 17:20 | Session 4, chaired by Qing Zhou, Zhejiang University |
| 15:30 - 16:00 | Asian Carp genomes provide insights into invasions and hybridization Guoqing Lu, University of Nebraska at Omaha, USA |
| 16:00 - 16:30 | Mutational signatures and selection pressures estimation in the cancer genome Zhixi Su, Fudan University |
| 16:30 - 17:00 | Three-Dimensional Genomics and Cancer [Abstract]
Cheng Li, Peking University
随着三维基因组技术的快速发展以及它的广泛应用前景,美国国立卫生研究院在2014年制定了4D Nucleome计划,从三维空间和时间尺度上研究细胞核内染色质的组织结构和功能。
我们研究组基于对癌症基因组中非整倍体变异频繁出现的原因和后果的研究兴趣,通过Hi-C实验和分析流程,研究多发性骨髓瘤细胞中非整倍体变异对三维基因组和表达谱的影响。
本报告将介绍三维基因组学背景以及我们组相关分析算法、数据库网站、癌症研究的进展。
|
| 17:00 - 17:20 | Genomics, Proteomics & Bioinformatics (GPB) — a rising journal in the field
Yuxia Jiao, Genomics Proteomics Bioinformatics |
| 18:00 - 20:00 | Welcome Dinner |
| October 13: Talks | |
| 09:00 - 10:10 | Session 5, chaired by Zhang Zhang, BIG, CAS |
| 09:00 - 09:40 | The BIG Data Center's Resources
Yiming Bao, BIG Data Center, BIG, CAS |
| 09:40 - 10:10 | Big data in psychiatric disorders for genetic study
Suhua Chang, Institute of Mental Health, Peking University |
| 10:10 - 10:30 | Tea & Coffee Break |
| 10:30 - 12:10 | Session 6, chaired by Wenming Zhao, BIG, CAS |
| 10:30 - 11:10 | Genetic diversity of long-tailed macaques (Macaca fascicularis) in Thailand: Application for biomedical research [Abstract]
Suchinda Malaivijitnond, Chulalongkorn University, Thailand
Long-tailed macaques (Macaca fascicularis) are one of the commonly used non-human primate (NHP) models for biomedical research and are the most encountered NHP species in Thailand.
Among 10 subspecies, common (M. f. fascicularis; Mff) and Burmese (M. f. aurea; Mfa) long-tailed macaques are found in Thailand.
Although primatologists denoted that genetics of Thai Mff are a mess because they carry the genetic admixture of rhesus macaques (M. mulatta; Mm).
Using 40 autosomal SNPs, the genetic admixture between Mff and Mm was maximized at the proposed hybrid zone (15-20 N) and the Mm gene flow to Mff population declined gradually in proportion to the distance from the hybrid zone.
Thenceforth, a Mff population at 16° 51′N contains 50% of Mm ancestry while a southern one living at 7° 12′N contains 15% of Mm ancestry. Turning a mess into opportunity, Thai Mff should be excellent animal model for drug and vaccine developments over other conspecific.
For example, Mm are susceptible to Plasmodium cynomolgi (a sister taxon of human P. vivax) and tuberculosis while Mff are tolerance to those diseases. Thus, Thai Mff who carry different degrees of Mm’s genetics should be varied in P. cynomolgi or tuberculosis infection.
Mfa are the only Asian NHP who can use stone tools to forage for encased foods. Exploring their genetics using partial mtDNA, Y-chromosome, whole mtDNA and whole genome sequences together with phylogenetic analysis, they are distinctive from their neighboring Mff species.
Regarding these unique characteristics, we proposed that their genetics have major effect on learning behavior and memory capacity. Thus, we are opening up the cognitive genes in this animal. Understanding their genes should be beneficial for neuroscience research while aging population is growing across the globe and the neurodegenerative diseases are increasing.
|
| 11:10 - 11:40 | Data Driven Biomedical Discovery [Abstract]
Hui Lu, Shanghai Jiao Tong University [Personal Profile]
Dr. Hui Lu is a Distinguished Professor and Head of the Department of Bioinformatics and Biostatistics, Shanghai Jiaotong University, the co-director of SJTU-Yale Joint Center for Biostatistics, and an adjunct professor of biostatistics of Yale University.
He is also the Director of Center for Biomedical Informatics in Shanghai Children’s Hospital. He got Bachelor’s degree from Peking University and PhD from University of Illinois at Urbana-Champaign. After that, he joined University of Illinois at Chicago as a faculty in Bioinformatics and got tenure in 2008.
His research areas include big data analysis in biomedical research, bioinformatics, biostatistics, molecular network modeling, systems biology, and drug design.
His current research interests are in translational medicine: integrating multi-omics data and disease phenotypes, constructing disease network, investigating self-adapting methods for clinical trial, set up high speed genomics data processing pipeline, large scale patient record analysis, building a phenotype-genotype based diagnosis system.
In recent years, data driven approaches have made a variety of contributions to disease analysis combining public data with in-house data in an attempt to increase biomedical understanding.
Popular topics include the discovery, prediction, and analysis of disease genes, statistical analysis of SNPs and disease, the prediction and discovery of new drug targets, the development of the disease ontology and its application to the human genome, the analysis of disease related molecular interaction networks, the development of “disease networks”.
In this talk we will present our effort in these areas. Specific examples include: combining genomics and clinical data in cancer biomarker discovery; joint analysis of genomic and imaging features; phenotype driven disease diagnostics; and building disease network using genomic data warehouse.
In summary, data driven approach has been playing key roles in translational medicine.
|
| 11:40 - 12:00 | EWAS Atlas: A knowledgebase of epigenome-wide association studies
Mengwei Li, BIG Data Center, BIG, CAS |
| 12:00 - 13:30 | Lunch |
| 13:30 - 15:10 | Session 7, chaired by Xiangfeng Wang, China Agricultural University |
| 13:30 - 14:10 | HOCOMOCO: towards the exhaustive inventory of binding motifs for human transcription factors [Abstract]
Vsevolod J. Makeev, Vavilov Institute of General Genetics, RAS, Russia
The abundant data on DNA-protein interactions for the first time make feasible to approach to an exhaustive inventory of binding motifs, possibly recognized by human transcription factors.
In our approach, we mostly used the ChIP-seq data, that appears to be most informative on the specificities of TF binding in vivo. We used five thousand ChIP-Seq experiments as the raw data,
the experimental datasets were uniformly processed within the BioUML framework using several ChIP-Seq peak calling tools and aggregated in the GTRD database.
As a result, we arrived at the new version of our HOCOMOCO (HOmo sapiens COmprehensive MOdel Collection), the curated collection of position weight matrix (PWM) models for binding sites for 680 human and 453 mouse TFs.
ChIPMunk software was used for systematic motif discovery. Motifs that displayed the best separation of the test (independent subsets of ChIP-Seq peaks) and emulated control datasets were selected for each transcription factor.
To reduce the number of irrelevant motifs emerged due to indirect binding we performed extensive computer assessment and human curation.
The current version of HOCOMOCO (v.11) includes 1302 mononucleotide and 576 dinucleotide position weight matrices, which describe primary binding motifs of each transcription factor and reliable alternative binding specificities. An interactive interface and bulk downloads are available on the web: http://hocomoco.autosome.ru and http://www.cbrc.kaust.edu.sa/hocomoco11. In my presentation, I will also discuss services that allow using HOCOMOCO for practical analyses in the fields of genetics and systems biology and the prospective of motif-based analysis of cell type specific binding of transcription factors.
|
| 14:10 - 14:40 | The sea cucumber genome provides insights into morphological evolution and visceral regeneration [Abstract]
Xiaojun Zhang, Institute of Oceanology, CAS
Apart from sharing common ancestry with chordates, sea cucumbers exhibit a unique morphology and exceptional regenerative capacity.
Here we present the complete genome sequence of an economically important sea cucumber, Apostichopus japonicus, generated using Illumina and PacBio platforms, to achieve an assembly of ~805 Mb (contig N50 of 190 Kb and scaffold N50 of 486 Kb), with 30,350 protein-coding genes and high continuity.
We used this resource to explore key genetic mechanisms behind the unique biological characters of sea cucumbers. Phylogenetic and comparative genomic analyses revealed the presence of marker genes associated with notochord and gill slits, suggesting that these chordate features were present in ancestral echinoderms.
The unique shape and weak mineralization of the sea cucumber adult body were also preliminarily explained by the contraction of biomineralization genes. Genome, transcriptome and proteome analyses of organ regrowth after inducted evisceration provided insight into the molecular underpinnings of visceral regeneration, including a specific tandem duplicated PSP94-like gene family, a significantly expanded FREP gene family and a positively selected Wnt signaling pathway.
This high-quality genome resource will provide a useful framework for future research into biological processes and evolution in deuterostomes, including remarkable regenerative abilities that could have medical applications.
Moreover, the multi-omics data will be of prime value for commercial sea cucumber breeding programs.
|
| 14:40 - 15:10 | A Gold Standard to Deconvolute Complicated Structures of Plant Genomes with Recursive Polyploidizations [Abstract]
Xiyin Wang, North China University of Science and Technology
Plants often have complex genomes, due to recursive polyploidizations and genome repatterning. This makes it difficult to deconvolute their genome structures,
and barrier the understanding their formation and the exploration of gene functional evolution. It would be a great pity if failing to decipher a newly sequenced genome structure when enormous amount of money and time invested.
However, such failures occurred quite often in last several years. Here, we propose a gold standard streamline to perform the genome structural analysis, adopted by quite several plant genome sequencing efforts,
which we suggest be taken as a gold-standard to analyze a new genome sequence. Using the streamline, we found an overlooked tetraploidization in the common ancestor of cucurbiteceae, which might have contributed to the fast divergence and establishment of the important family of plants.
|
| 15:10 - 15:30 | Tea & Coffee Break |
| 15:30 - 17:10 | Session 8, chaired by Zhixi Su, Fudan University |
| 15:30 - 16:10 | Darkness in the human gene and protein function space: Modest illumination by the life science literature, the trend for protein function discovery decline since 2000 and the importance of medical record databases [Abstract]
Frank Eisenhaber, Bioinformatics Institute, Singapore [Personal Profile]
Frank Eisenhaber studied mathematics at the Humboldt-University in Berlin and biophysics and
medicine at the Pirogov Medical University in Moscow. He was awarded a MD in 1985. Three years
later, he received the PhD in molecular biology from the Engelhardt Institute of Molecular Biology
in Moscow (supervision by Dr. Vladimir Gayevich Tumanyan). After postdoctoral work at the
Institute of Molecular Biology in Berlin-Buch (1989-1991) and at the EMBL in Heidelberg (1991-
1999), he worked as teamleader of the bioinformatics research group and head of the general IT
department at the Institute of Molecular Pathology (IMP) in Vienna (1999-2007). He joined the
Bioinformatics Institute A*STAR Singapore in August 2007 and he is the Executive Director since
August 2013.
It is generally believed that full human genome sequencing was a watershed event in human history that boosted biomedical research, biomolecular mechanism discovery and life science applications.
At the same time, researchers in the field of genome annotation see that there is a persisting, substantial body of functionally insufficiently or completely not characterized genes (for example, ~10,000 protein-coding in the human genome) despite the availability of full genome sequences.
A survey of the biomedical literature shows that the number of reported new protein functions had been steadily growing until 2000 but the trend reversed to a dramatic decline thereafter (1,2). The fastest growing set of genes in the last decade is that with 500 or more full publication equivalents, i.e., the genes that are well characterized anyhow.
At the same time, the annual amount of life science publications doubled between 2000 and 2017. There are no apparent scientific or financial reasons for the decline in biomolecular mechanism discovery; probably, current instruments of science funding do not direct or even discourage researchers to go after the difficult problems of gene function discovery.
The talk will cover examples of protein-coding gene function discovery in the GPI lipid anchor biosynthesis pathway (3,4), details about the role of natural organism research (5) and medical patient databases (6).
|
| 16:10 - 16:40 | Factors, pitfalls and solutions for genomic selection in crops [Abstract]
Xiangfeng Wang, China Agricultural University
Genomic selection (GS) has been successfully applied in cattle breeding. Considering unique aspects in crop breeding, how GS can be efficiently and properly used in crops still requires sufficient practices with real-world data, such as to solve prediction of hybrid vigor performance for crops like maize.
With a breeding population consisting 9,873 F1 individuals representing 23% of complete crosses of 1428 female and 30 male lines covering the six heterosis groups in maize, we first evaluated general factors that may influence GS accuracies.
These factors include proportion of training and testing set, marker density and marker type, population stratification, trait heritability and so on. Then, we discuss pitfalls covering multiple aspects that are general mistakes need to avoid when applying GS.
These pitfalls include ways of phenotypic data processing, GWAS-based marker selection, improper training frameworks biasing model evaluation, factors usually neglected in a GS model, and so on.
Last, we raise an optimal framework for designing a GS scheme to reach maximum predictability and objectiveness, and propose a possible solution for nominating combinations with potential yield heterosis based on the “omnigenic model” hypothesis using machine learning strategies.
|
| 16:40 - 17:10 | Biocloud platform enhances bioinformatics studies in the big data era
Hongkun Zheng, Biomarker Technologies |
| October 14: Talks and BHBD | |
| 09:00 - 10:10 | Session 9, chaired by Zhang Zhang, BIG, CAS |
| 09:00 - 09:40 |
Small genomes, big data: Improving bacterial genome annotation one COG at a time [Abstract]
Michael Y. Galperin, National Center for Biotechnology Information, USA [Personal Profile]
Dr. Michael Y. Galperin has received his PhD from the Lomonosov Moscow State University in Russia and postdoctoral training at the University of Louisville and University of Connecticut.
He has been at the NCBI Computational Biology Branch since 1996, first as a GenBank Fellow, then Staff Scientist, and currently as a Lead Scientist. He has published more than 200 research papers, reviews and book chapters and is an author of a textbook on comparative genomics.
In 2008-2017, he served as the editor of the Nucleic Acids Research annual Database Issue. He currently serves as an editor of the Genomics Updates section in Environmental Microbiology and, since July 1st 2018, an editor of the Journal of Bacteriology.
Microbial genome sequencing projects all over the world continue to flood public databases with sequences of deduced proteins, only a small fraction of which has been ever studied experimentally or could be studied in detail any time soon.
The only feasible way to assign funcions to these proteins is to tentatively predict them through computational analysis. The Clusters of Orthologous Genes (COG) database, first created in 1997, has been a popular tool for functional annotation.
Its success was largely based on (i) its reliance on complete microbial genomes, which allowed unequivocal assignment of orthologs and paralogs; (ii) family-based approach, which used the functions of the characterized members of the protein family (COG) to assign function to the entire family and describe the range of the potential functions when there were more than one, and (iii) careful manual curation of the COG names and protein contents.
The latest update of the COG database allowed to take an unbiased view at the progress in genome annotation and evaluate the problems and challenges in assigning functions to the remaining uncharacterized and poorly characterized open reading frames.
Combining functional data from UniProt, RefSeq, Pfam, InterPro, and CDD databases, as well as the experimental data from original papers kept in PubMed and PubMed Central showed that original COG assignments had error rate of < 0.5%. Many tentative COG predictions have now been verified, either by direct experiments or through high-throughput methods. Functional assignments have been made for some widespread conserved proteins, many of which turned out to participate in translation, including rRNA maturation, tRNA modification, and similar processes.
From the practical point of view, COGs can be useful for quality assurance of genomic sequences and for the identification of (i) “holes” in metabolic pathways and functional systems; (ii) unique enzymes that might be used as drug targets, and (iii) conserved genes and operons coding for previously overlooked functional systems.
|
| 09:40 - 10:10 | Chemoresistance in Patients with Acute Leukemia: from Molecular Targets to Clonal Evolution [Abstract]
Qianfei Wang, BIG, CAS
Major therapeutic progress using cytotoxic agent has been made in leukemia over the last fifty years, yet chemoresistance remains an unmet clinical challenge.
Approximate 10-25% of newly diagnosed AML and most relapsed patients have a poor drug response and low rate of long-term survival.
Although advances in cancer genomics has greatly increased our understanding of the molecular characteristics in tumor biology, recent studies suggest that Darwinian evolution of intratumor heterogeneity represent a major challenge to develop therapeutic strategy for improved disease control.
I will present our recent genomic and functional studies involving the JAK-STAT pathway and the methyl-transferase SETD2, as well as mutational landscape in refractory and relapsed AML under sequential treatment of induction regimens. I will discuss the limitations of molecular targeting and how evolutionary principles can be applied during the treatment of leukemia.
|
| 10:10 - 10:30 | Tea & Coffee Break |
| 10:30 - 11:40 | BHBD Session, chaired by Yiming Bao, BHBD Coordinator |
| 10:30 - 10:40 | Welcome Speech & Introductory Remarks |
| 10:40 - 11:00 | Introduction to Global Biodiversity & Health Big Data Alliance (BHBD) |
| 11:00 - 11:10 | Speeches by Senior Officials and IUBS Representative |
| 11:10 - 11:20 | Speeches by BHBD Founding Member Representatives |
| 11:20 - 11:30 | Opening Ceremony |
| 11:30 - 11:40 | Group Photo |
| 12:00 - 14:00 | Lunch |
| 14:00 - 16:00 | The 1st BHBD Alliance Meeting (invited only), 1104 at the 11st floor |
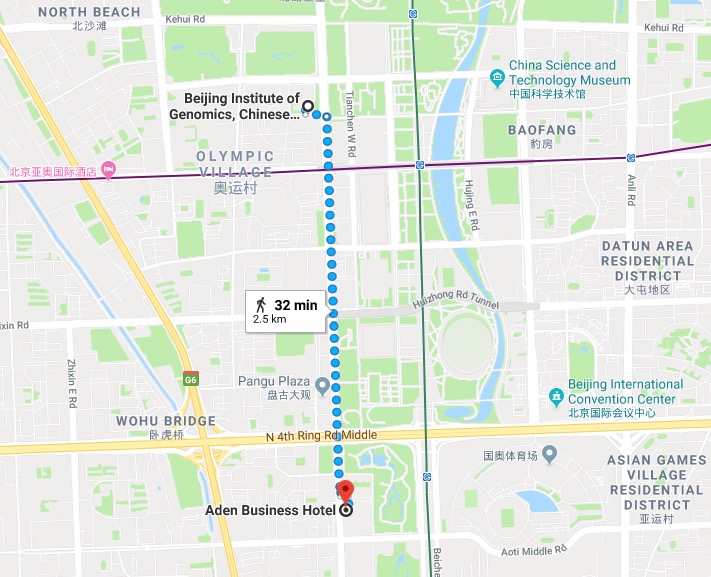
Location
Hotel
-
Aden Business Hotel (北京亚丁湾商务酒店)
9 Minzuyuan Road, Chaoyang District
Beijing 100101, China
Tel: +86 (10) 5936-0606 -